The continuing bioprocess automation brings new challenges. Industry 4.0 pushes us to strengthen the cohesion between people, processes and technologies. How can we develop systems and functions that provide an enhanced user experience and enable a substantial, organic relationship between software and people? Tacking on new functionality to existing solutions is one option, but the opportunity to rebuild using proven new approaches is too important to miss. Here we take you behind the scenes of our software development.

INFORS HT bioprocess platform eve® will be used as an example of one way of thinking about meeting existing and future needs for bioprocesses. As examples will be specific, the principles are general and apply widely. The key factors you will find in this overview cover:
- Choice of database structure, i.e. new approaches to store and combine data
- The user interface for the “smartphone generation”
- Accommodation third-party devices, e.g. a range of bioreactors and
- Interface with other software .e.g. modelling software
- Having data on one platform, in one place, but accessible to many disparate groups
- Scalability – from a single user with one device to a company-wide solution
1. A NoSQL database structure for big data
This is the prime example of a new technology being applied because of a fresh start. Bioprocess software solutions are traditionally based on Structured Query Language (SQL) databases. SQL databases primarily hold information on process values, setpoint, etc. A key thing to know about SQL databases is that they are relational. Their structure depends on tables of data in rows and columns, with relationships between tables defined in a schema. The schema is fixed, and adding new data fields is challenging. Also, a relational database is not ideal for storing data in document formats or from a wide variety of sources. Finally, scaling a relational database is very resource-intensive.
This led to the concept of NoSQL (Not Only SQL) databases being developed in the last decade. They are easily scalable and handle many types of data as “documents” (in the sense of individual records of information). These documents can be searched by using structured language. An important difference to many SQL solutions is that a NoSQL database will normally be tightly integrated into a software solution for a wider purpose, such as bioprocess control. Data is stored as a series of indices.
Many of the good features of SQL databases have been added to NoSQL solutions, such as the ability to be transactional, i.e. only making entries when the “transaction” is complete, such as starting or stopping a batch. NoSQL databases have been good choices for historian data solutions, as they can handle large amounts of disparate data quickly.
Forty years ago, bioprocess data would not have met all these criteria, as sensor inputs were more limited and data for quality control as well as set up would be stored elsewhere than in the main SCADA (Supervisory Control And Data Acquisition) software. Now the emphasis is shifting towards statistical models of bioprocess control and large data sets are becoming the norm.
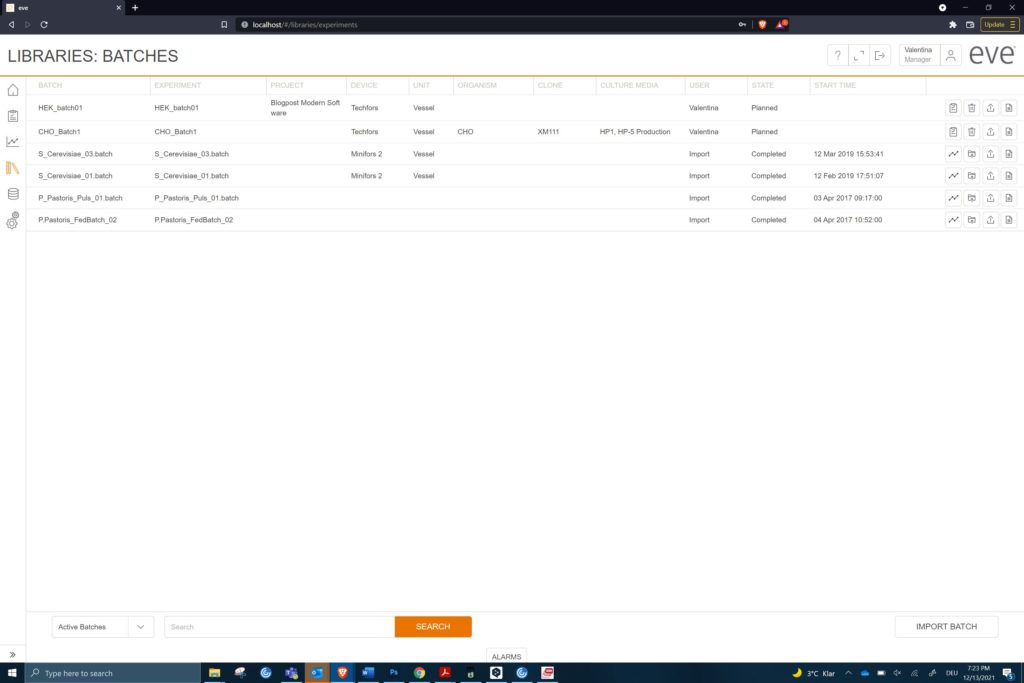
Access to the data is primarily from the bioprocess software due to tight integration in software solutions (c) Infors AG 2021
2. A Browser-based User Interface (UI)
A feature of many of the legacy SCADA programs for bioprocess control is the use of a Microsoft Office® style of interface. Tabs, ribbons with icons, and information in both the top and bottom borders of resizable windows is the paradigm. This is familiar and easy to navigate for most users. However, as more interaction with software takes place via the internet, the question is whether this style of interface is so comfortable for people used to living and working from their browser? Again, the most common way of doing something has changed over time, and the user interface (UI) of modern software needed to adapt.
The interactive features of a browser interface represent a simplification over multiple ribbons and icons. As an interface to access specific software, other advantages are apparent:
- It is not tied to the main software. The software needs only to be updated in one place for all users to take advantage of the change. No roll-out over multiple computers.
- The user interface is mostly browser-agnostic, as a set of features common to all are used.
- Mobile devices have browsers and can also access the software. More work in laboratories is being done on tablet-sized devices, so this gives all approved users easy access no matter where their location is on-site.
- If allowed, remote access over the internet is possible. Integral security features and good IT policies can make this a secure, convenient way of checking a running system remotely. This is separate from the ability to send alarm messages by email if things are not going according to plan
- Even the computer running the bioprocess software accesses its UI via a web browser, so a common interface is enforced. No training or distinction between local and remote users.
Bioprocess software UI in a browser windows (c) Infors AG 2021
3. Open to third party hardware
The range of devices that could be connected to bioprocess software has multiplied since the early days of SCADA running on a PC. This trend is growing, and new devices are also bringing a more data-rich environment to shake flasks, bags and even micro-titre plates. The trend of data exchangers over a network has taken hold in the last decade. Protocols such as OPC (Open Process Control) in several flavours are common amongst manufacturers.
Devices can be connected to bioreactors in several ways, many of them simple and still valuable after decades of use i.e.
- Analogue signals, usually 4-20mA or 0-10V
- Modbus serial connection for low-level communications
- Serial outputs, occasionally requiring conversion to a network protocol
This is not usually done by the bioprocess software, but these extra values can expand the needs for data exchange from a basic ten parameters to over thirty. Add in analytical data and calculated soft sensors and the total can be much higher.
Modern bioprocess software must integrate hardware from some or all the following categories:
These types of hardware can be mixed and multiplied for many environments, from research laboratories through pilot halls to production plants. Even with common protocols, different generations of device firmware often require drivers to be created for reliable data transfer. All of this needs a reliable base of code on which to overlay the specifics. That can be best achieved when there are no legacy considerations.
The ability to accommodate all the extras must be a feature of modern bioprocess software. The need to understand the biology behind the process is taking over. This marks a shift from simple optimisation of physical parameters, such as temperature.
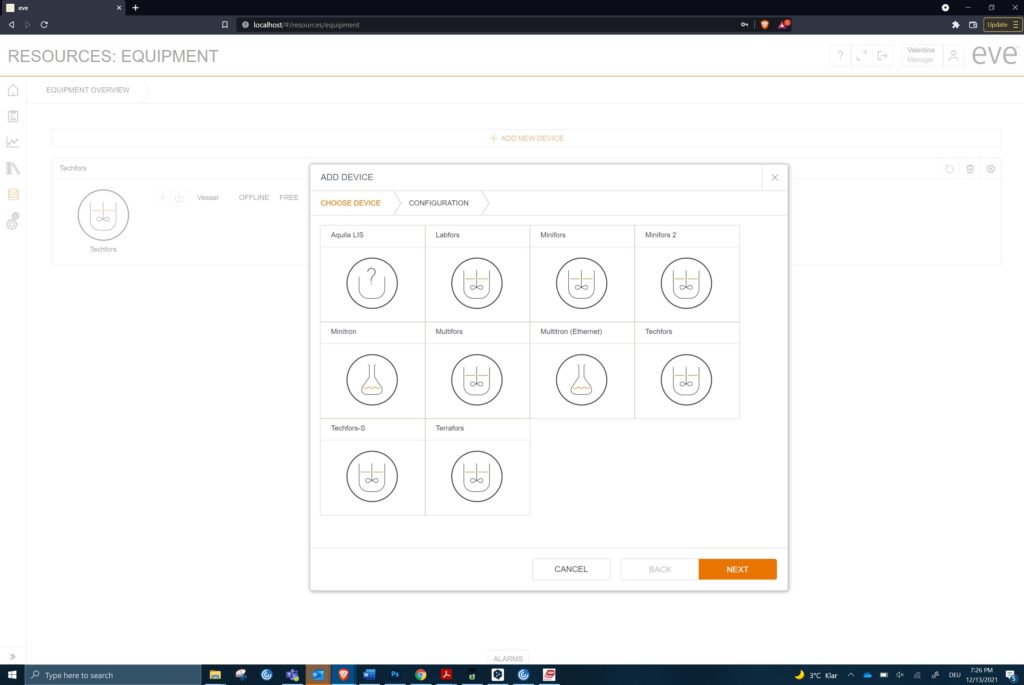
Equipment from just one manufacturer can generate many choices (c) Infors AG 2021
4. Open to third party software
This is the corresponding story to link diverse hardware. The ability to import and export data has been a feature of SCADA bioprocess software for decades. This can be the manual entry of offline analytical data, export of values as Comma-Separated Values (CSV) into spreadsheets or real-time data exchange via long-established protocols such as Object Linking and Embedding (OLE).
There is a good, modern alternative, based on the technologies of the internet. The Representational State Transfer Application Program Interface (RESTful API) provides a consistent interface for software to communicate over the web. It uses common web protocols to give access to remote data using simple commands.
RESTful API allows:
- Data from one or more batches to be exported from the bioprocess software
- External software, e.g. a modelling package, can access the data
- Processed data and its values could be sent back to the bioprocess software and used in real-time
Some work is needed to set up the necessary commands. However, this needs only simple scripting rather than coding.
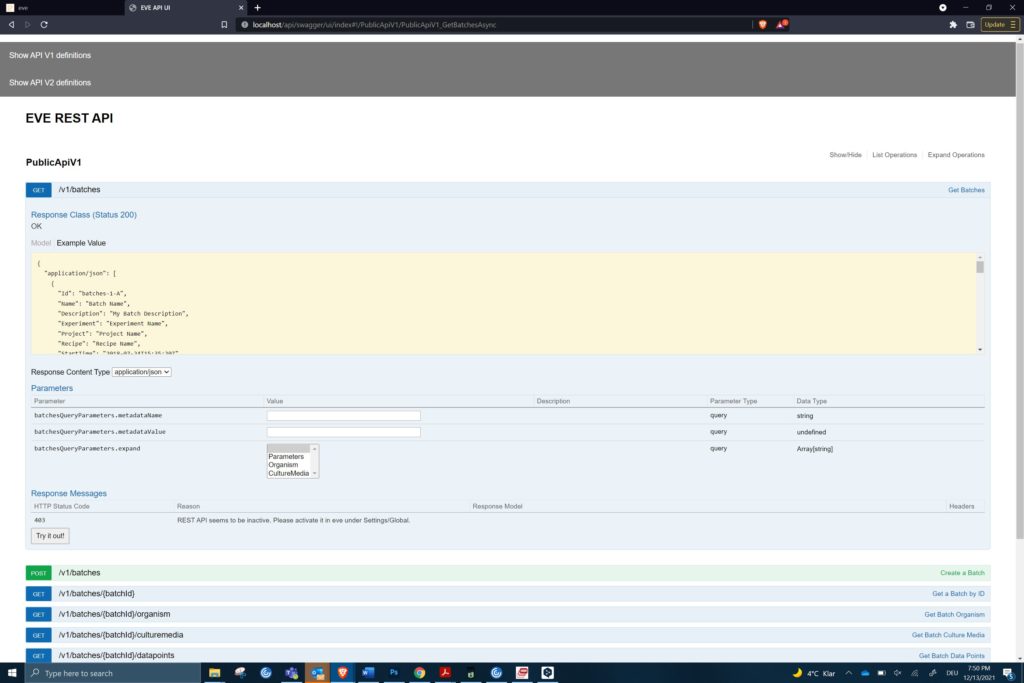
An example of the REST API to get batch data from eve® (c) Infors AG 2021
5. All your data in one place and accessible to many
SCADA-based bioprocess software is involved with several types of data, i.e. process values, set-points alarm and time. Of course, configuration and recipe data are part of the software, but may not be accessible in reports. What it has not handled so well is data with different formats and which is based on documents rather than critical values. It can include audit trail data, user management and project planning aspects. In the past, these would be handled by different groups, departments and individuals with a variety of software solutions (or paper forms). This approach allows data to spread throughout an organisation and following it can be time-consuming and costly.
The people involved in preparing, running and analysing the results of a batch process have also changed. Quality control and external regulatory authorities need access to archived data. Managers need to know which experiments of a set have been completed and which are yet to be started. Optimising the allocation of resources can depend on this data being easily available. That means being accessible to all these special interest groups in real-time and without disturbing running processes.
Keeping all the data in one place is a good reason to use a NoSQL database. Different sorts of documents store the data, but can be interrogated into tailored reports. Access via a web browser means no group must have special software installed to get at the data they require. Of course, this has relevance to larger companies and institutions, but the same data structure is there even for a single user.
The types of user also need to be defined in groups, with each group having full access to what they need but not the entire range of capabilities. So, an administrator will differ from a manager. A guest user, e.g. a regulatory inspector, can access batch data and audit trails but not interfere with a running process. The ability to enforce strong passwords with ageing helps with secure access. Individual log-ins and passwords can identify each user separately. This information appears on each entry on the audit trail.
The question of how necessary this is to each user depends on the institution, application and oversight of the bioprocess. A central repository of data accessed through a single interface has attractions in terms of security, training requirements and the preparations needed for regulatory inspections.
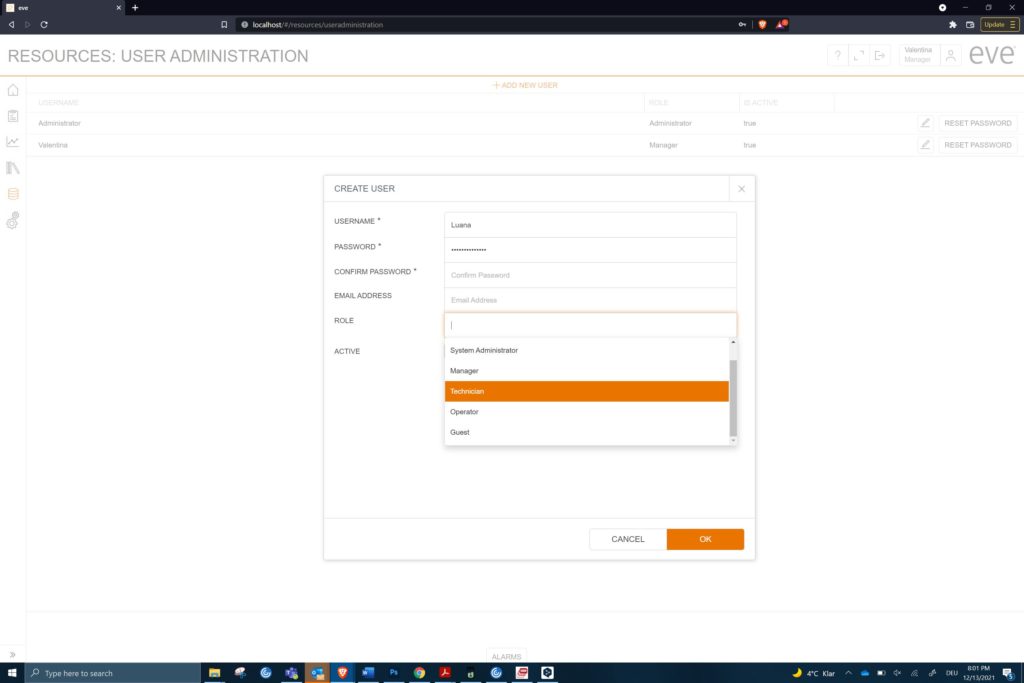
Different users cover a wide range of permissions and capabilities. (c) Infors AG 2021
6. Scalability
SCADA systems have often been created with multiple devices in mind. At a low level, this involves adding connections and setting up a new hardware configuration. What is not so easy is scaling for the ever-increasing quantity of data. SQL databases need ever more resources to cope and a limit can soon be reached. A NoSQL database provides a way to bypass the issue of handling big data. It can store information, index it and make it available quickly on any scale.
How does that help a single user with just one or two connected items of equipment? By providing the potential to grow without the need to learn new software or ways of doing things. Even a solo user can benefit from the storage of disparate types of data, quick access via a browser, the tools for analysis and workflow control. The aim is to succeed, and that will bring new equipment, more colleagues and greater contact with pilot and production facilities. That is when a scalable system can remove a lot of potential problems.
There may be a need for new computing hardware and an upgrade of the operating system, but these are low-cost items. This may be done using a Virtual Machine (VM) running on a local or remote service. So long as a fast Ethernet connection (Gigabit) is available to the equipment, this is a suitable solution for larger institutions.
Keeping an established, familiar system can save a lot of time and money. With nothing more than an IP address and login details, others can be included, making expansion effortless. The ability to join other groups and share data across sites worldwide comes from the way the software is structured, even in the simplest package. It’s about limiting yourself to what you need and thinking about how to make future expansion as easy as possible.
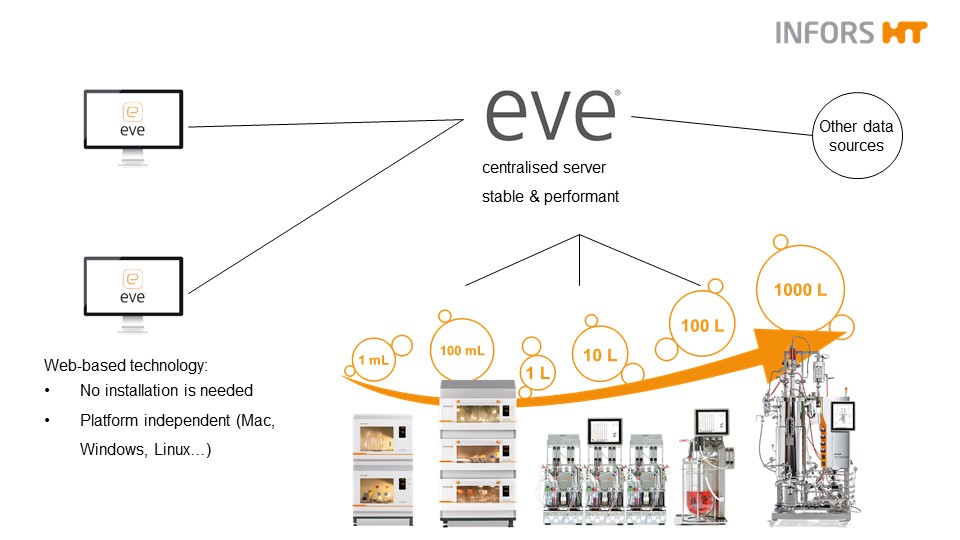
One software solution from smallest to largest networks (c) Infors AG 2021
Summary
This overview describes how an opportunity to use modern technologies for the creation of bioprocess software can provide a stepping stone to the use of big data, bioprocess automation and access by many to a single repository of diverse types of data. A simple web browser provides the user interface and keeps pace with the changing face of software. A scalable solution for data storage ensures high future-proofing, as automation using process modelling takes bioprocessing to the next level. An outward-facing approach to linking both hardware and software allows one software solution to work in many situations.
To come back to the initial question, this may interest you if
- Your data volume and velocity are increasing rapidly
- You need a scalable solution for many users across different bioprocess hardware and software
- You want a common interface that needs no installation and maintenance
- You wish to keep valuable data in a single location but with good access
An overview can only use a broad-brush approach. However, if it helps in thinking about what modern bioprocess software needs to do for you, that’s a plus.
Any relevant comments, ideas for new posts and constructive criticism are always welcome.
Thanks for taking the time to read this. Tony Allman, INFORS HT
